|
2019-03-03 14:42:47 -0600
| received badge | ● Popular Question
(source)
|
|
2018-03-19 23:28:03 -0600
| received badge | ● Notable Question
(source)
|
|
2016-04-18 11:40:53 -0600
| received badge | ● Popular Question
(source)
|
|
2014-05-31 10:39:37 -0600
| commented question | Qt5 + OpenCV 2.4.9 cv::imshow error I don't recall if you must have Qt installed. Do you have it? |
|
2014-05-31 10:32:39 -0600
| commented question | HOGDescriptor returning wrong result in static image What is the foundWeights's value for those windows? Your threshold might be too low. Try to increase it, test in this picture and in a picture with real persons to see if you won't lose anybody. |
|
2014-05-28 14:35:09 -0600
| received badge | ● Self-Learner
(source)
|
|
2014-05-28 14:10:29 -0600
| answered a question | Gaussian Kernel with arbitrary size m x n One may take advantage of kernel separatibility to generate such Gaussian Kernel. // mu: mean, sigma: standard deviation
float gaussianDistribution(float x, float mu, float sigma) {
return std::exp( -(((x-mu)/(sigma))*((x-mu)/(sigma)))/2.0 ) / (cv::sqrt(2 * CV_PI) * sigma);
}
cv::Mat_<float> getGaussianMask(int nrows, int ncols) {
cv::Mat_<float> gaussianMask = Mat_<float>::zeros(nrows, ncols);
// compute a bidimensional gaussian
for (int x = 0; x < gaussianMask.cols; ++x) {
for (int y = 0; y < gaussianMask.rows; ++y) {
// taking advantage of the gaussian's kernel separatibility
gaussianMask(y,x) =
gaussianDistribution(y, gaussianMask.rows/2 - 1, gaussianMask.rows/4 - 1) *
gaussianDistribution(x, gaussianMask.cols/2 - 1, gaussianMask.cols/4 - 1);
}
}
// normalizing
cv::Scalar totalsum;
totalsum = cv::sum(cv::sum(gaussianMask));
gaussianMask = gaussianMask.mul(1.0/totalsum[0]);
}
Resulting image for nrows = 128 and ncols = 64. 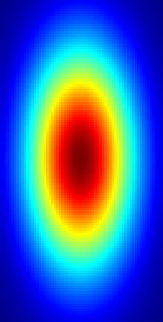
You may also vary the size of the standard deviation sigma to produce Gaussians with smaller or bigger radius. Here, I'm using gaussianMask.rows/4 and gaussianMask.cols/4. The mean mu determines the center of the Gaussian. // ...
gaussianDistribution(y, gaussianMask.rows/2 - 1, gaussianMask.rows/4 - 1) *
gaussianDistribution(x, gaussianMask.cols/2 - 1, gaussianMask.cols/4 - 1);
I used cv::applyColorMap(gaussianMask, gaussianMask, cv::COLORMAP_JET); to get this coloring. |
|
2014-05-27 19:26:48 -0600
| commented answer | Retrieve HOG Detector scores Thank you, Siegfried! I totally missed that. It is not a documented feature, though :) |
|
2014-05-26 15:42:30 -0600
| asked a question | Retrieve HOG Detector scores I'm using the default HOG Detector to detect pedestrians, such as in the code below. HOGDescriptor hog;
hog.setSVMDetector(HOGDescriptor::getDefaultPeopleDetector());
vector<Rect> found, found_filtered;
hog.detectMultiScale(data, found, 0, Size(strideX,strideY), Size(32,32), scale, 2);
I want to retrieve the score that the classifier assigned to each sample. For instance, found[0] has 0.9 confidence to be a pedestrian; found[1] has 1.3 confidence score; and so on. I want these because I will use them to make a plot missrate vs false positives per image (FPPI). Is there any way to retrieve the classifier responses for each sample? |
|
2014-05-21 15:08:07 -0600
| asked a question | Gaussian Kernel with arbitrary size m x n Is it possible to generate a Gaussian kernel with an arbitrary size m x n? How can I do that?
I've looked into getGaussianKernel, however, it returns only squared kernels. In case I need to implement it by hand, any suggestions are welcome. |
|
2014-02-12 07:37:03 -0600
| received badge | ● Nice Question
(source)
|
|
2014-01-13 12:03:07 -0600
| commented answer | Training a soft cascade classifier Hey, @StevenPuttemans! I have one question about this paper. I thought that the Soft Cascade by itself provided a speedup of 20 times (Table 1 of the paper). The impression it gives is that by using only Soft Cascade we already can achieve this 20x speed up. Using the stixels and ground plane assumptions, we can increase this speed up even further. Is that correct? I mean, shouldn't the speed up provided by the Soft Cascade be independent of the other optimization approaches? |
|
2013-12-19 10:34:21 -0600
| received badge | ● Self-Learner
(source)
|
|
2013-12-19 10:34:21 -0600
| received badge | ● Necromancer
(source)
|
|
2013-12-19 10:24:20 -0600
| answered a question | How do I remove duplicate rows in cv::Mat? This is a solution without STL. It is very ugly, though. // keep only unique rows
Mat allPairs; // Mat with duplicate values
Mat uniqueStrides; // Mat that will contain the unique values
uniqueStrides.push_back( allPairs.row(0) );
for (int i = 1; i < allPairs.rows; ++i) {
int isInside = false;
for (int j = 0; j < uniqueStrides.rows; ++j) {
int count = 0;
for (int k = 0; k < uniqueStrides.cols; ++k) // checks by element of
if(allPairs.at<int>(i,k) == uniqueStrides.at<int>(j,k))
++count;
if (count == 2) {
isInside = true;
break;
}
}
if (isInside == false) uniqueStrides.push_back( allPairs.row(i) );
}
This algorithm checks for every uniqueStrides entry and for every value. As my matrices are n x 2, I check if the number of equal values is two: ...
if (count == 2)
isInside = true;
...
If positive, then this row is already in my matrix. |
|
2013-12-19 10:01:36 -0600
| commented answer | How do I remove duplicate rows in cv::Mat? berak and I came to a conclusion that this approach might not work for every input. Depending on the order of a and b, the result may return true or false for the same pair of Mats. We considered to take the absolute value of the sum, however this solution does not respect Strick Weak Ordering. I think it is not possible to remove duplicates using STL when your matrix has positive and negative values. But, this answer still very helpful because it can work for other input and explains how to implement comparators for STL. |
|
2013-12-19 07:43:57 -0600
| commented answer | Why Kalman Filter keeps returning the same prediction? @Siegfried, I think my question has changed. But, instead of updating my question, I'll accept your answer because it solves this problem and ask my new question (the one from the comment above) in another post. What do you think? |
|
2013-12-14 16:52:50 -0600
| commented answer | Why Kalman Filter keeps returning the same prediction? What happens if I track without new observations? If I set a non-zero velocity and give the initial position of a pedestrian, then I can track this person without new observations, assuming that the velocity is constant. If I feed Kalman with new observations given by the pedestrian detector, hence I'm using tracking by detection, which is more accurate because it can correct Kalman if a pedestrian changes its direction. Is this correct? |
|
2013-12-02 14:42:44 -0600
| received badge | ● Student
(source)
|
|
2013-12-02 09:27:03 -0600
| received badge | ● Editor
(source)
|
|
2013-12-02 09:26:29 -0600
| asked a question | Why Kalman Filter keeps returning the same prediction? I'm trying to track pedestrians on a video using Kalman Filter. First, I initialize Kalman with the top-left coordinate (x0,y0) of a pedestrian's bounding box, which is stored within a sample. Next, I call kalmanPredict(), but it keeps returning the same (x0,y0) coordinate I passed. for (int i = 0; i < samples->size(); i++) {
// retrieves person
sample = samples->at(i);
// Gets the bounding box of a person
Rect rectSample = sample->GetWindow();
// x0 and y0 coordinates of the bounding box
Point2f aux(rectSample.x0,rectSample.y0);
// initializes Kalman with this coordinate
initKalman(aux.x,aux.y);
// predicts for 10 frames, for example
for (int j = 0; j < 10; j++) {
//Predict
p = kalmanPredict();
s = kalmanCorrect(p.x,p.y);
//crop image and send for shared memory
SSFRect ret(p.x,p.y,rectSample.w,rectSample.h);
Point pt1, pt2, pt3;
pt1.x = p.x;
pt1.y = p.y;
pt2.x = p.x + rectSample.w;
pt2.y = p.y + rectSample.h;
pt3.x = s.x;
pt3.y = s.y;
rectangle(data, pt1, pt2, cv::Scalar(0,255,0), 5);
rectangle(data, pt3, pt2, cv::Scalar(255,0,0), 1);
// show result
imshow("janela", data);
waitKey(30);
aux = pt1;
cout<<i;
}
}
The following code is the setup of Kalman I'm using and contains the functions kalmanPredict() and kalmanCorrect(). I have tried different values for KF.transitionMatrix and KF.measurementMatrix, but it didn't make difference. void initKalman(float x, float y)
{
// Instantate Kalman Filter with
// 4 dynamic parameters and 2 measurement parameters,
// where my measurement is: 2D location of object,
// and dynamic is: 2D location and 2D velocity.
KF.init(4, 2, 0);
measurement = Mat_<float>::zeros(2,1);
measurement.at<float>(0, 0) = x;
measurement.at<float>(1, 0) = y;
KF.statePre.setTo(0);
KF.statePre.at<float>(0, 0) = x;
KF.statePre.at<float>(1, 0) = y;
KF.statePost.setTo(0);
KF.statePost.at<float>(0, 0) = x;
KF.statePost.at<float>(1, 0) = y;
KF.transitionMatrix = (Mat_<float>(4, 4) << 1,0,1,0, 0,1,0,1, 0,0,1,0, 0,0,0,1); // Including velocity
KF.measurementMatrix = (Mat_<float>(2, 4) << 1,0,0,0, 0,1,0,0);
setIdentity(KF.processNoiseCov, Scalar::all(.005)); //adjust this for faster convergence - but higher noise
setIdentity(KF.measurementNoiseCov, Scalar::all(1e-1));
setIdentity(KF.errorCovPost, Scalar::all(.1));
}
Point kalmanPredict()
{
Mat prediction = KF.predict();
Point predictPt(prediction.at<float>(0),prediction.at<float>(1));
/*This prevents the filter from getting stuck in a state when no corrections are executed.*/
KF.statePre.copyTo(KF.statePost);
KF.errorCovPre.copyTo(KF.errorCovPost);
return predictPt;
}
Point2f kalmanCorrect(float x, float y)
{
measurement(0) = x;
measurement(1) = y;
Mat estimated = KF.correct(measurement);
Point2f statePt(estimated.at<float>(0),estimated.at<float>(1));
return statePt;
}
What am I missing? Why Kalman keeps returning the same prediction value to me? |
|
2013-10-22 15:04:38 -0600
| commented question | C++ Face Detection |
|
2013-10-15 10:39:44 -0600
| marked best answer | How do I remove duplicate rows in cv::Mat? I have four n x 2 matrices and I want to concatenate them, but removing the duplicate rows. I tried to use std::set std::set<cv::Mat> nonDuplicatedPairs;
for (int i = 0; i < drPairs.rows; ++i) {
nonDuplicatedPairs.insert( deltasMat.row(i) );
nonDuplicatedPairs.insert( drPairs.row(i) );
nonDuplicatedPairs.insert( dlPairs.row(i) );
nonDuplicatedPairs.insert( ulPairs.row(i) );
}
but it gives the following compilation error: error C2440: 'return' : cannot convert from 'cv::MatExpr' to 'bool' These matrices are of type CV_32SC and have only one channel. |